TS-PScan
Pan-cancer screening tool. Classifies H&E whole-slide images as cancer or normal across 11 tissue types. Outputs a calibrated risk score and attention heatmap. Available for research access upon request.
Tsebix develops AI models for cancer detection from H&E whole-slide images.
A cancer diagnosis depends on a pathologist reading a slide, often at 40× magnification, across thousands of tiles, under relentless case volume. Subspecialty expertise is scarce everywhere: demand is outpacing the workforce in every major healthcare system. The diagnostic gap is not a geography problem. It is a scale problem.
Tsebix develops AI models that work alongside pathologists, surfacing regions of concern and scoring malignancy likelihood from H&E whole-slide images. No additional staining or on-premise hardware required.
Pan-cancer screening tool. Classifies H&E whole-slide images as cancer or normal across 11 tissue types. Outputs a calibrated risk score and attention heatmap. Available for research access upon request.
Breast cancer analysis. Detects invasive carcinoma and classifies histological subtype (IDC vs ILC). Predicts ER, PR and HER2 status from H&E images.
Lung cancer analysis. Differentiates adenocarcinoma from squamous cell carcinoma from H&E whole-slide images.
Colorectal cancer analysis. Cancer detection and histological subtype classification from H&E whole-slide images
Whole-slide images are orders of magnitude larger than natural images, and the signal is sparse: a diagnosis can rest on a single cluster of cells among tens of thousands of tiles. Our pipeline combines foundation-model tile embeddings with attention-based aggregation, trained under weak supervision from slide-level labels — not hand-drawn pixel masks.
Model performance varies across institutions due to differences in scanners and staining protocols. Tsebix addresses this through a site-specific calibration step performed during onboarding — no retraining required.
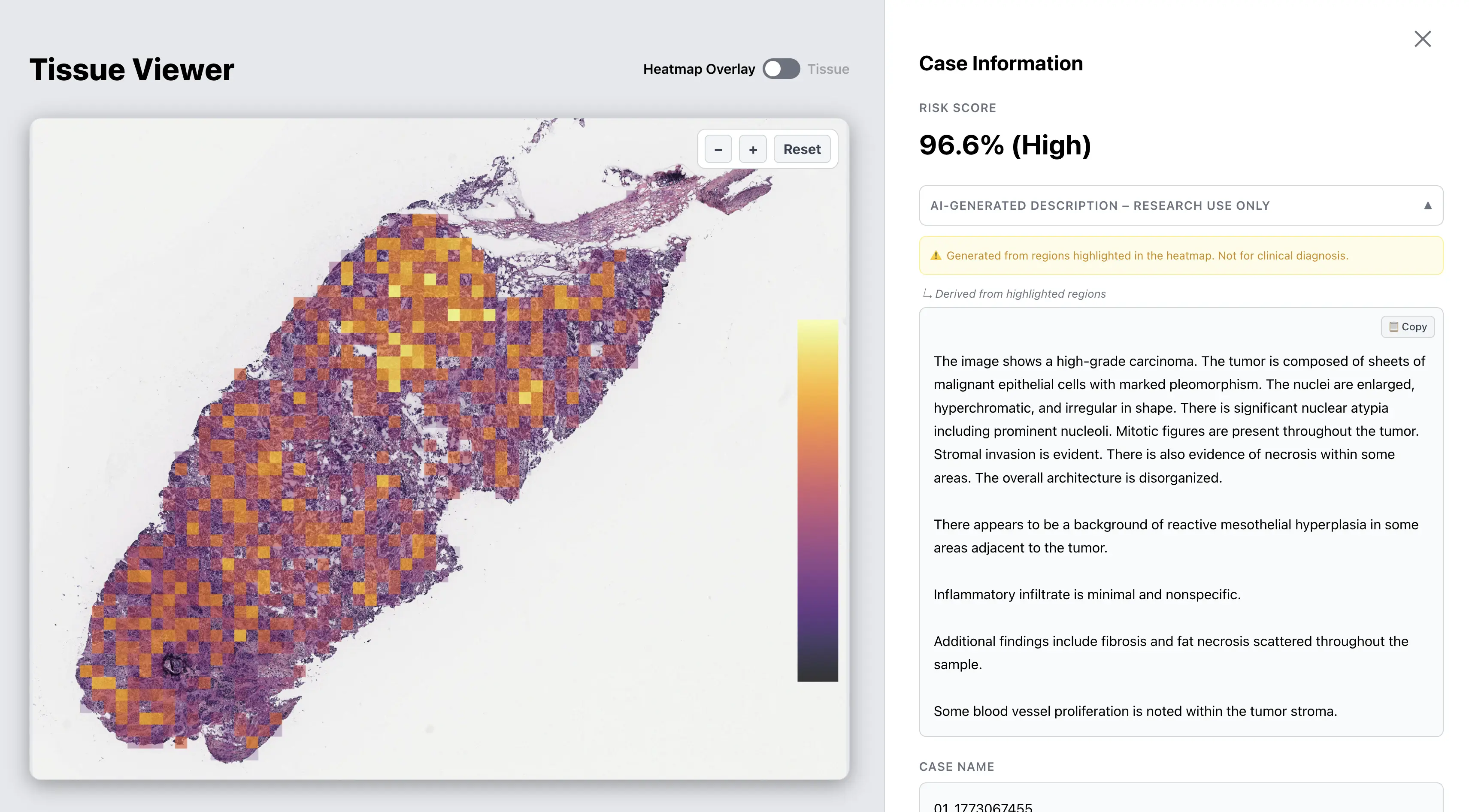
Trained on TCGA + GTEx. Independently tested on CPTAC and CMB across 11 cancer types.
TS-Breast predicts key pathological features directly from H&E. Models for PR and HER2 status prediction are currently in active validation.
TS-Lung differentiates primary lung cancer subtypes directly from H&E. Models for EGFR mutation status prediction are currently in active validation.
TS-Colon detects colorectal cancer and classifies histological subtype from H&E. Models for MSI status prediction are currently in active validation.
Team
Advisors
Tsebix products are currently available under research-use agreements with hospitals, academic groups and clinical study sponsors. Terms are scoped to the study.
Start a conversationWe review every inbound request. If there is a fit, we will share validation data, a scoped research agreement, and a timeline for integration.